Publications
2026
Genomics of host-microbiome interactions in humans
Nature Reviews Genetics, 2026
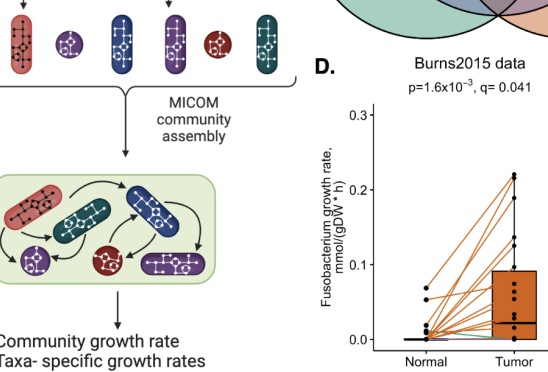
Metabolic modeling and functional genomics reveal taxa and host gene interactions in colorectal cancer
bioRxiv, 2026
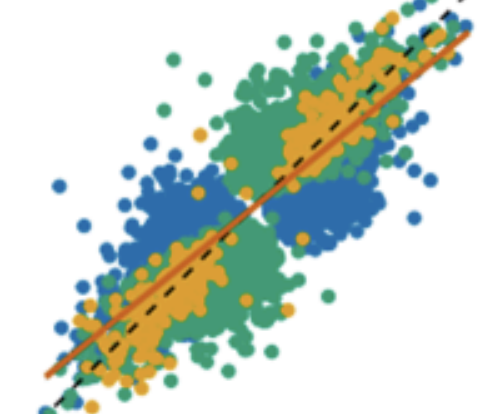
Host genotypes interact with microbial communities to modulate gene expression in the human intestine
medRxiv, 2026
2025
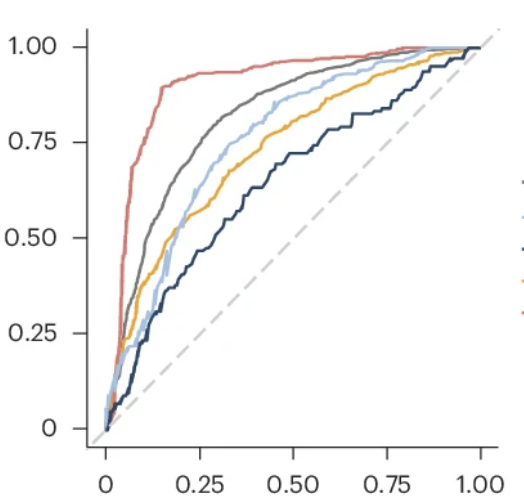
HIV and antiretroviral therapies exert distinct influences across diverse gut microbiomes
Nature Microbiology, 2025
2022
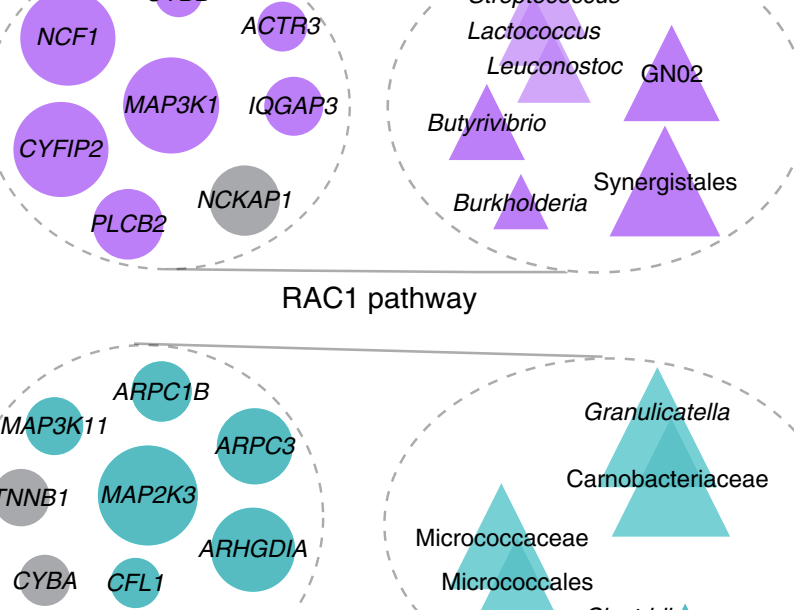
Identification of shared and disease-specific host gene–microbiome associations across human diseases using multi-omic integration
Nature Microbiology, 7, 780–795, 2022
2021
The Gut Microbiome in Konzo
Nature Communications, 2021
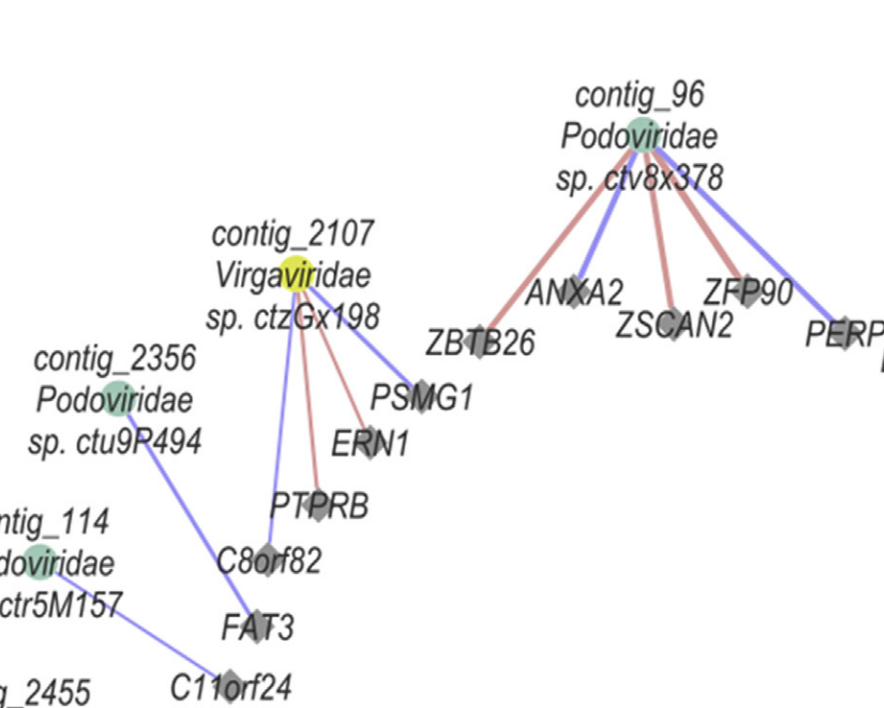
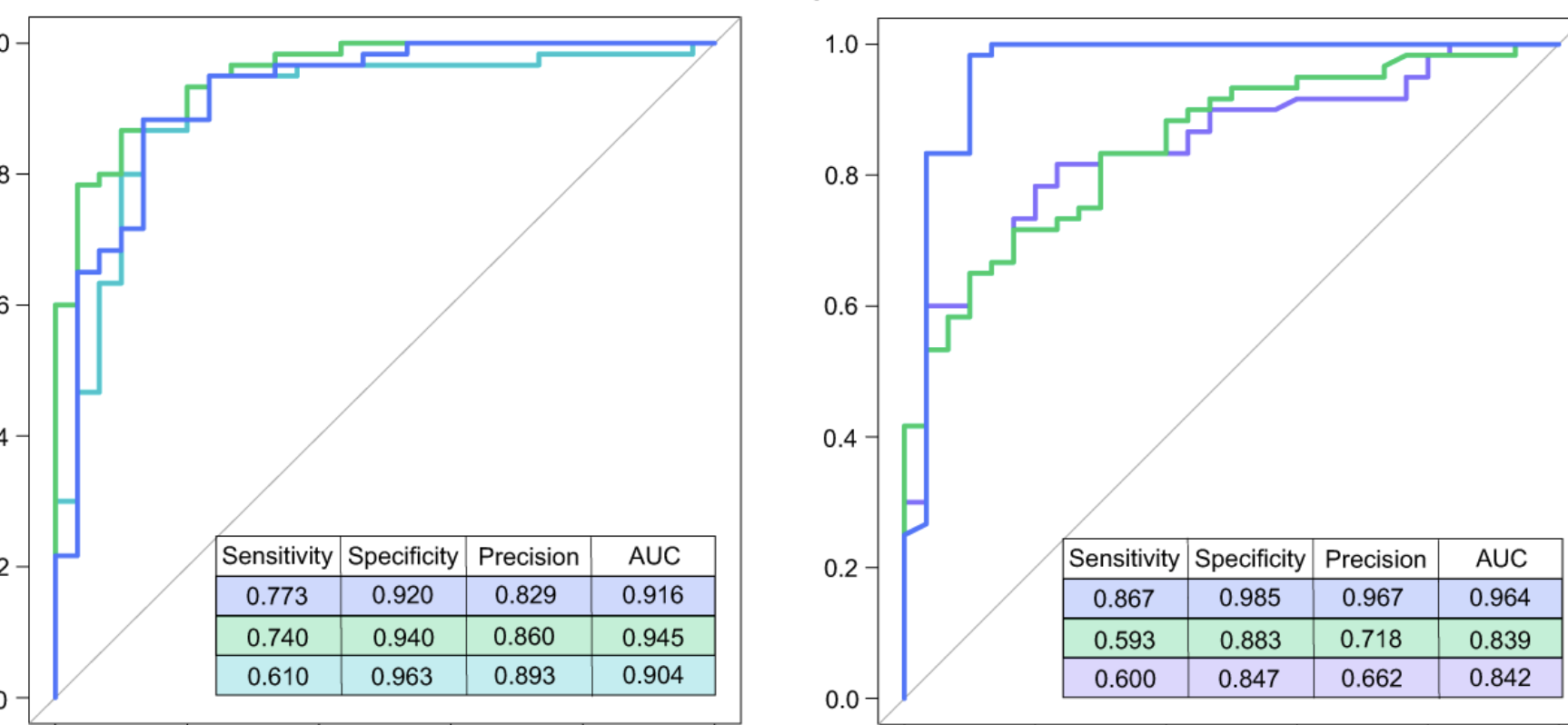
Multi-omics analyses show disease, diet, and transcriptome interactions with the virome
Gastroenterology, 2021
2020
Longitudinal Multi-omics Reveals Subset-Specific Mechanisms Underlying Irritable Bowel Syndrome
Cell, 182(6), 2020

Interactions between the gut microbiome and host gene regulation in cystic fibrosis
Genome Medicine, 12(12), 2020
2019
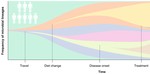

Population dynamics of the human gut microbiome: change is the only constant
Genome Biology, 20, 150, 2019
2018
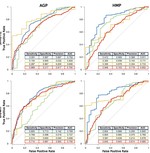

Gut microbiota diversity across ethnicities in the United States
PLoS Biology, 16(12), 2018
Distinct microbes, metabolites, and ecologies define the microbiome in deficient and proficient mismatch repair colorectal cancers
Genome Medicine, 10(78), 2018

Colorectal cancer mutational profiles correlate with defined microbial communities in the tumor microenvironment
PLoS Genetics, 14(6), 2018
Transposon mutagenesis screen in mice identifies TM9SF2 as a novel colorectal cancer oncogene
Scientific Reports, 2018
2017

HOMINID: A framework for identifying associations between host genetic variation and microbiome composition
GigaScience, 6(12), 2017
2015
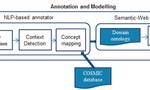
A Semantic Web-based System for Mining Genetic Mutations in Cancer Clinical Trials
AMIA Summits on Translational Science Proceedings, 142–146, 2015
2014

Exploring Linked Data with Contextual Tag Clouds
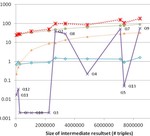
Partitioning OWL Knowledge Bases for Parallel Reasoning
IEEE International Conference on Semantic Computing, 108–115, 2014
2013
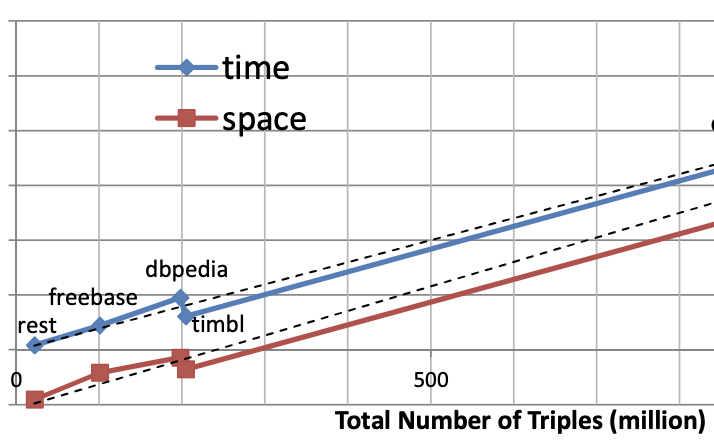
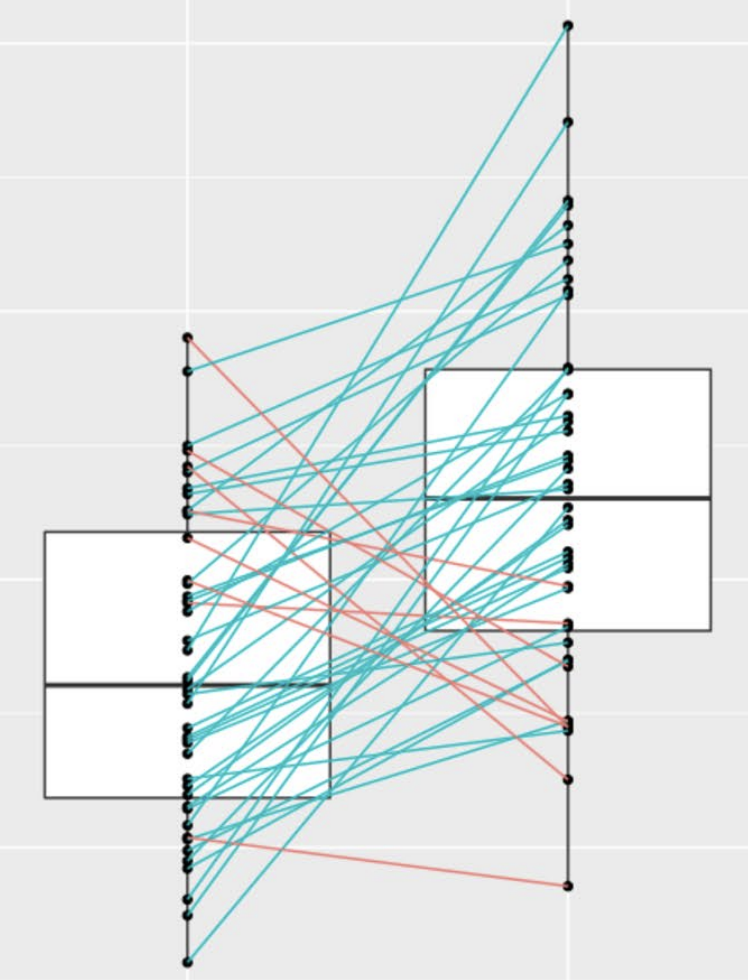
Infrastructure for Efficient Exploration of Large Scale Linked Data via Contextual Tag Clouds
The Semantic Web – ISWC 2013, Lecture Notes in Computer Science, vol 8218, 2013